Neuroscience Applications
Neuroscience Applications
Build a Comprehensive Whole-Brain Atlas with Integrated Multiomics
Accurately characterizing every individual cell within the brain is challenging due to the low expression of many functionally important genes. Using MERFISH 2.0 Technology and InSituPlex® multiplex immunofluorescence provides complementary spatial views of RNA and protein within the same tissue regions, connecting gene expression programs to protein phenotypes and revealing cell states, interactions, and pathway activities at single-cell resolution.
Neuroscience-focused gene panels include key markers for microglia, astrocytes, and oligodendrocytes, and can also functionally map immune cell infiltration in brain tumors. When used alongside protein panels, researchers can investigate molecular and cellular changes associated with aging and neurodegeneration.
MERFISH Mouse Brain Atlas
To demonstrate the power of the MERSCOPE® Platform, we generated the MERFISH Mouse Brain Receptor Map that includes canonical cell type markers, nonsensory G-Protein coupled receptors (GPCRs), and Receptor Tyrosine Kinases (RTKs).
We designed this specific gene panel because nonsensory GPCRs mediate signaling and may play vital roles in brain aging and neurodegenerative disorders. GPCRs are notoriously difficult to analyze, making the ability to completely spatially profile GPCR expression across the brain with cellular context particularly valuable.
MERFISH for Neuroscience Isn’t Just Mapping
Vizgen’s MERFISH 2.0 pre-designed neuroscience panels, the MERSCOPE™ Mouse Pan Neuro Panel, 815 Gene (v2.0), and the MERSCOPE™ Human Brain Panel, 815 Gene (v2.0) are ready-to-run, curated gene panels designed to map cell types and functional states across the mouse and human central nervous system on the MERSCOPE® and MERSCOPE Ultra™ Platforms.
PanNeuro Cell Type Panel V2 Capabilities
Whole-Brain Atlases
Generate comprehensive high-resolution cell-type maps across diverse brain regions.
Neuronal Subtypes & Circuits
Resolve rare cell types and map spatially organized neural pathways.
Activity-Dependent Genes
Capture immediate-early gene activation (e.g., c-Fos, Arc) after learning or stimulation.
Neurodevelopment & Aging
Profile molecular trajectories across development and aging.
Glial Biology & Neuroinflammation
Characterize astrocyte, oligodendrocyte, and microglial heterogeneity in health and disease.
Neurological Disease Models
Perform spatial profiling in Alzheimer’s, Parkinson’s, ALS, epilepsy, and other disorders.
Building a mouse brain atlas starts with the right markers. To address this need, we developed a 960-gene human brain MERFISH panel that enables deep spatial profiling of neuronal subtypes and Alzheimer’s-associated pathology using the MERSCOPE® Ultra Platform.
Insights Gained from MERSCOPE Data
Cellular Mapping of the Brain
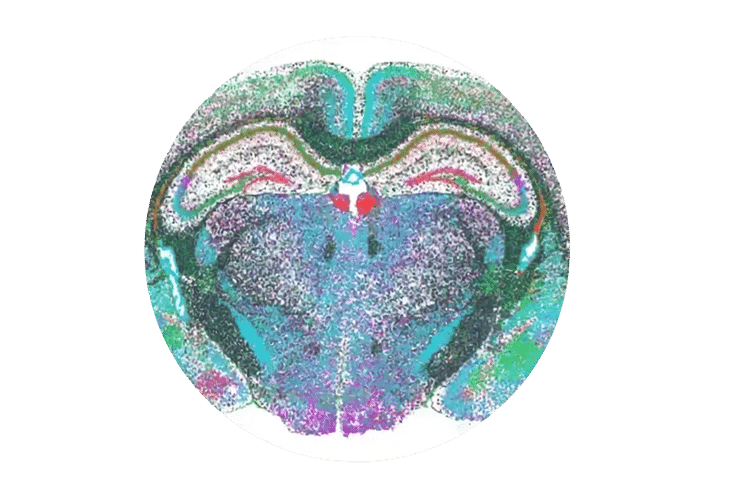
MERSCOPE’s high sensitivity allows us to map gene expression by cell type and see the variation in the gene expression pattern of receptors within the mouse brain.

Figure 3: The variation in the gene expression pattern of eight different receptors between neuronal and non-neuronal cell types and their positions within the mouse brain are detectable by MERSCOPE.
Aging Brain: Spatial “Clock” of Gene Expression
Researchers profiled mouse brains across adulthood, mapping cell types and age-related gene programs in coronal and sagittal sections to build a mouse brain atlas using the MERSCOPE Platform. The analysis shows shifts in cellular composition and spatially localized, age-responsive trajectories.
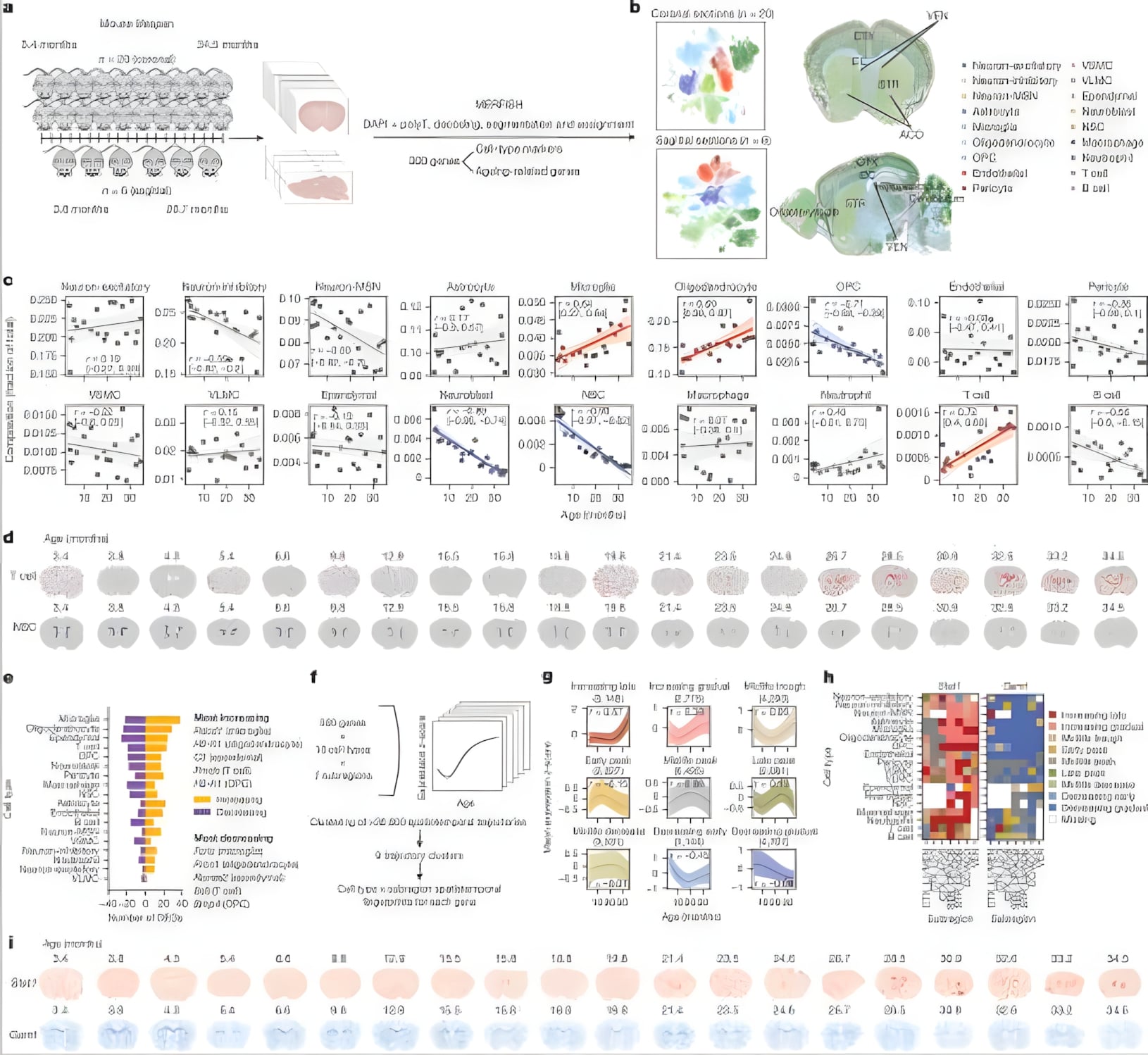

Figure 4: MERFISH cell maps and trend plots reveal age-linked changes in composition and gene expression (e.g., Stat1, Gamt) across brain regions. (Sun et al., Nature 2024)
Developing Cortex: Cell–Cell Communication
Researchers used MERFISH to define cell “niches” across six human neocortical samples and quantified neighborhood enrichment and directed signaling between cell types. Ligand–receptor analysis revealed key developmental pathways, including neuregulin and somatostatin signaling.
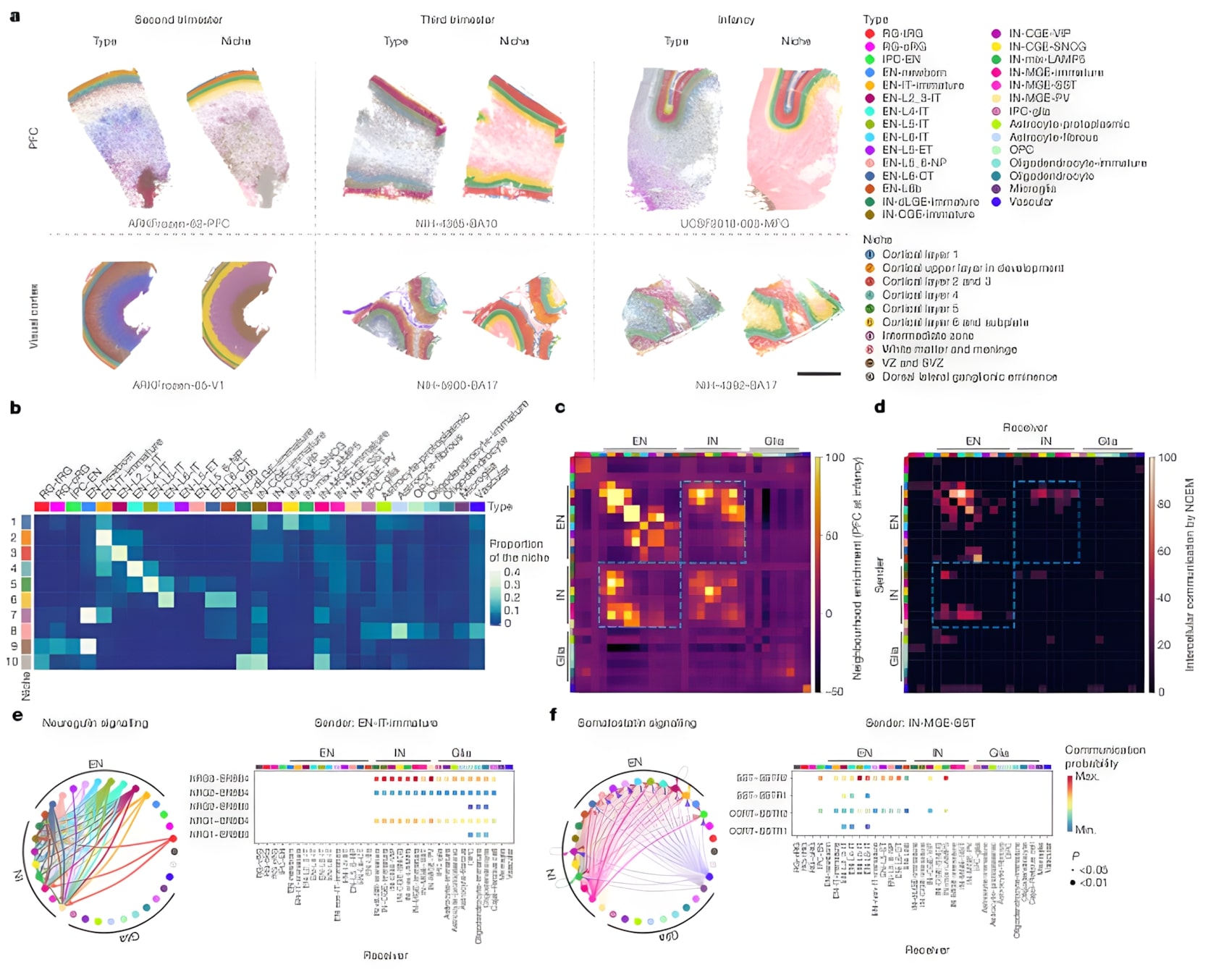
Figure 5: Spatial niche maps with matrices of enrichment and intercellular communication; directional signaling probabilities for NRG and SST pathways. (Wang et al., Nature 2025)
Publications
The MERSCOPE® and MERSCOPE® Ultra Platforms are widely used in neuroscience research, including for building whole-brain atlases, because they generate true spatial transcriptomic data across entire tissue sections down to the subcellular level. With MERFISH 2.0, MERSCOPE Ultra enables detailed cellular and functional mapping of the brain, providing deeper insights into neurological processes and disease mechanisms.