The importance of understanding the relationship between cells & their locations inside tissues
Why is spatial context important to understanding human health? When we think about how biological systems work, they are incredibly complex. To study these intricate systems, we simplify them by focusing on a smaller area– a particular organ or cell type – usually in an artificial environment such as a petri dish or tissue culture flask. However, such static measurements don’t always provide the necessary insight into the mechanisms at play under natural conditions.
Cells and tissues exist in a three-dimensional space where communication networks and specific spatial arrangement are critical to proper function. Spatial genomics provides us with this context, combining the unparalleled level of detail afforded by single-cell sequencing with the ability to map the three-dimensional position of each cell – or transcript with a cell – precisely. To give you some perspective in terms of numbers: Vizgen’s MERSCOPE® Platform can currently run panels of up to 500 genes, and look at around a million different cells within a tissue section, resulting in the ability to detect up to a billion RNA transcripts within that tissue section. Today’s technology is bringing us closer to creating a full picture of how cells behave in their natural state, surrounded by their neighbors and in a state of constant communication and adaptation to the changing environment. With this wealth of information, we can gather important insights into how cells adapt to change and what goes wrong under different disease conditions.
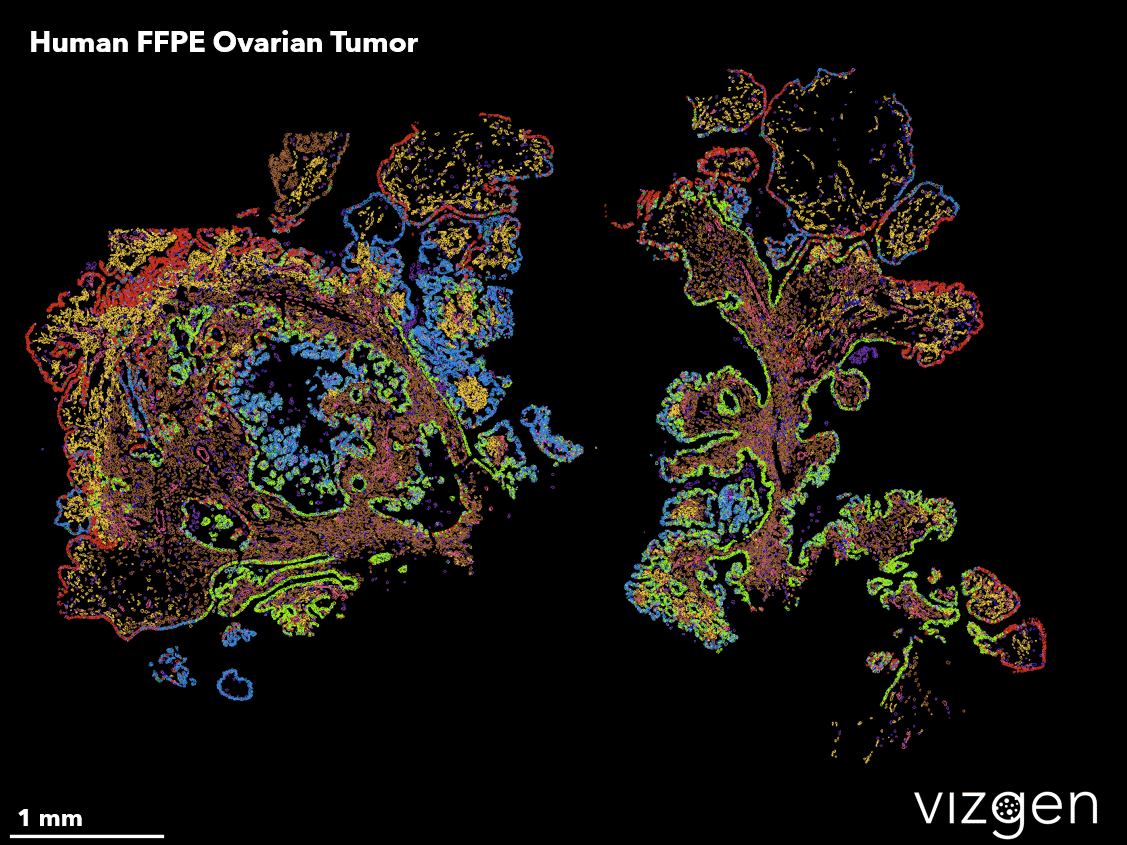
One of the ways spatial genomics is currently being leveraged to improve human health is in the study of cancer. A solid tumor is a complex and ever-changing unit containing multiple cell types that interact with both the tumor itself as well as healthy surrounding tissue. Because of its complexity and the difficulty of replicating the tumor microenvironment in a controlled lab setting, little is currently known about how these cells interact and the consequences of changes in their behavior. Spatial genomics allows researchers to study tumors in their natural state, providing valuable insight into how they adapt over time and respond to various therapies. This ability to track and characterize cell types will have important implications in predicting clinical outcomes, and providing more informed, specific care to cancer patients.
Spatially resolved transcriptomics was named the “Method of the Year” by Nature Methods in 2020, underscoring how this technology has grown in sophistication to allow incredibly detailed analysis of single cells while retaining the spatial context. As this technology continues to improve, we will see the spatial mapping of cells more readily applied to clinical situations. Going beyond just the transcriptome adds another layer of complexity and a better understanding of the systems at work. This vast amount of data and the identification of prognostic and diagnostic biomarkers will enable clinicians to provide better, more specific treatment for individual patients.