Introducing the MERSCOPE FFPE Anti-body Co-staining Data Release
We are thrilled to announce the launch of our Protein Co-Detection Data Release, demonstrating the capability of spatial multiomics profiling with our MERSCOPE® Platform to unravel immuno-oncology signatures in uterine cancer.
The data release combines MERFISH with protein co-staining for CD45RA and CD45RO, key immune cell markers involved in T-cell differentiation and activation, to provide a multi-dimensional view of the tumor microenvironment within human uterine Formalin-Fixed Paraffin-Embedded (FFPE) samples.
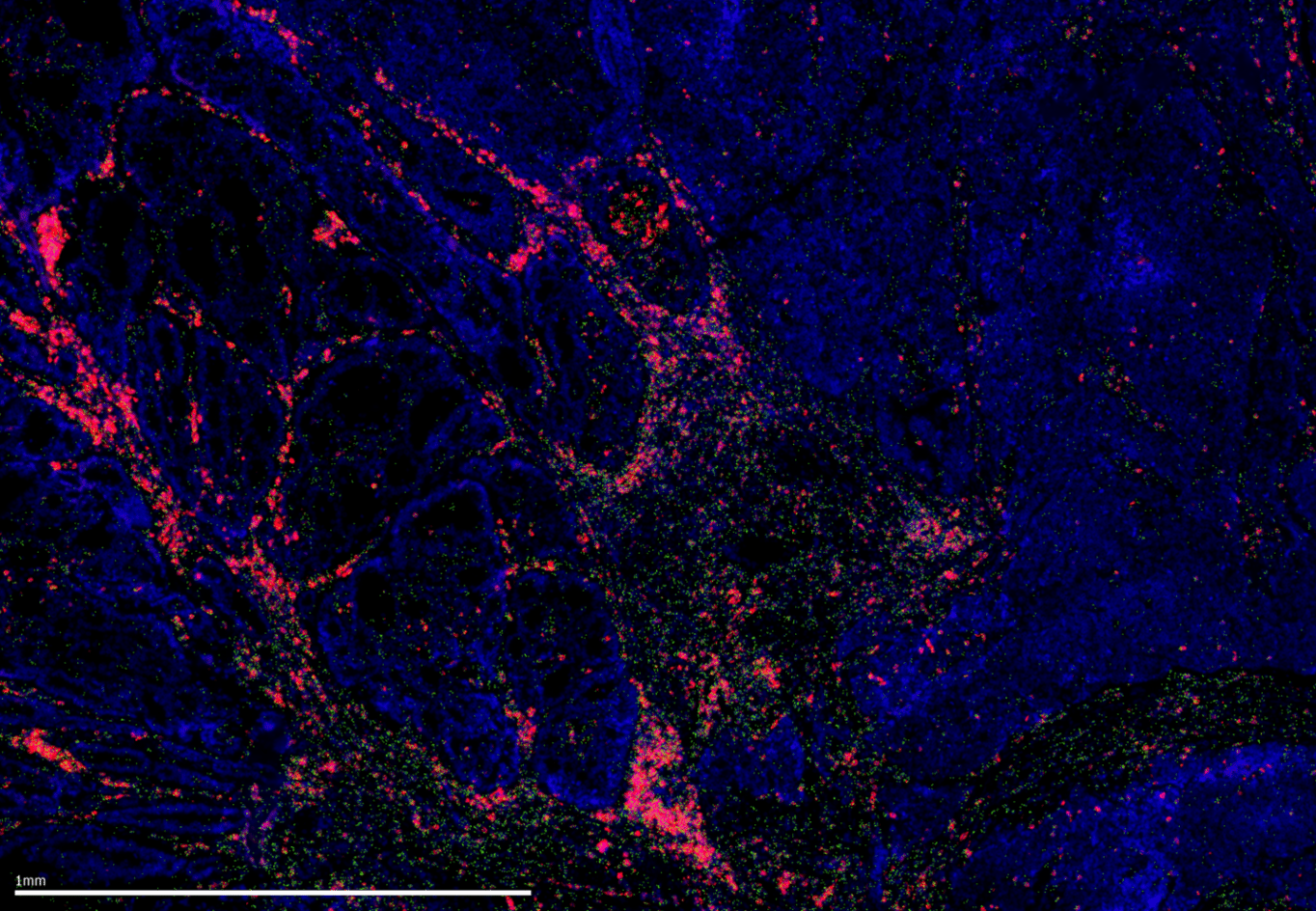
Figure 1. MERSCOPE® Platform Data Image showing the expression of PRTRC mRNA and CD45RO protein in FFPE human uterine cancer tissue.
Analyzing these multiomics datasets not only resolves different cell types with distinct transcriptomic features and their spatial organization/interaction in the tissue, but also enables the identification of specific immune cell type/gene expression patterns that correlate with the presence or absence of CD45RA and CD45RO. This provides a powerfully comprehensive picture of the immuno-oncology landscape in uterine cancer.
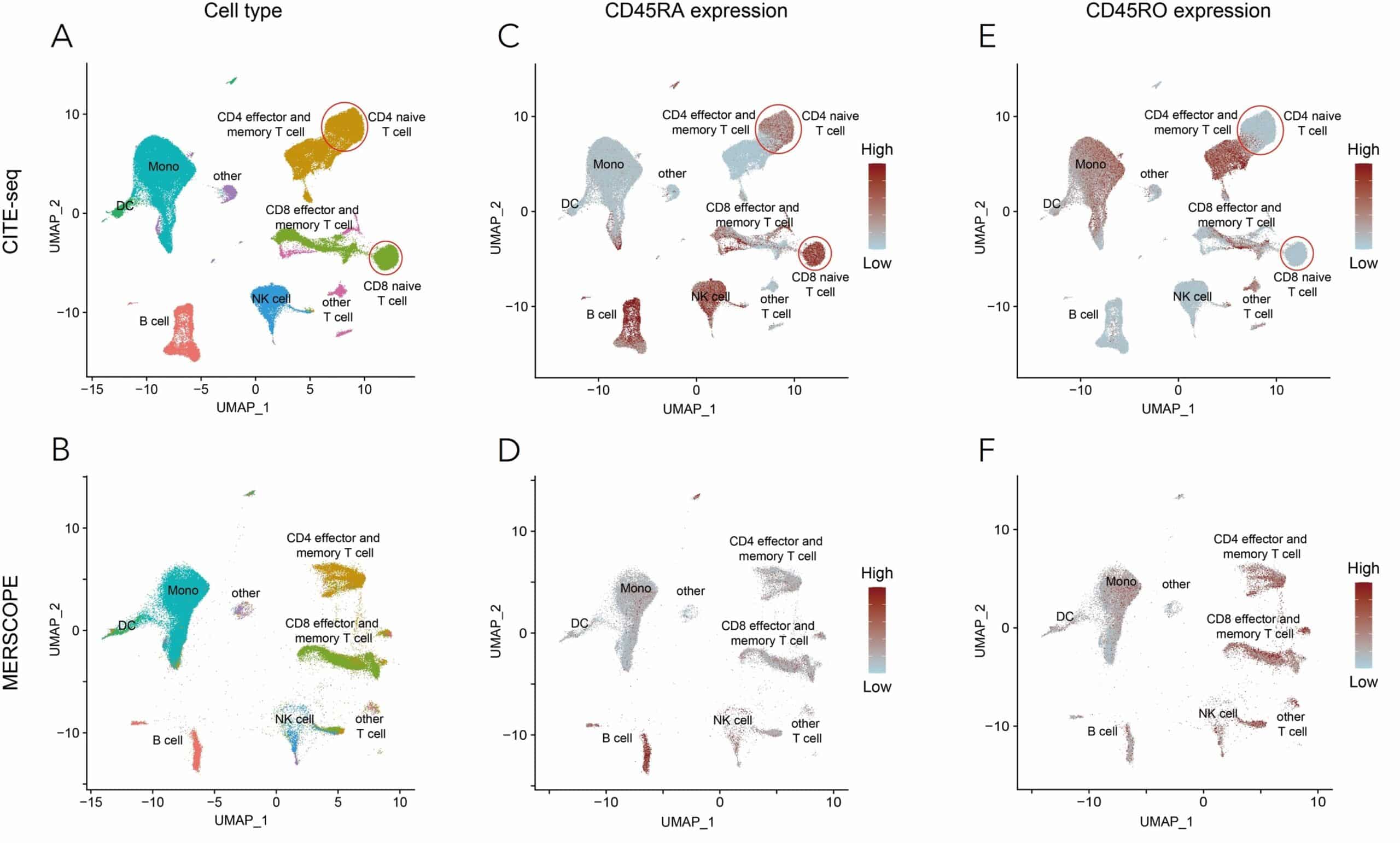
Figure 2. Accurate protein detection in human uterine cancer tissue to validate the results of protein co-detection, the immune cell types of MERSCOPE datasets were extracted and integrated with published CITE-seq data of PBMC. (A) UMAP visualization of immune cell types in CITEseq data. (B) UMAP visualization of immune cell types identified in uterine cancer MERSCOPE datasets. The cell type labels are predicted from integration with CITE-seq data, and cells are embedded in the same UMAP space as (A). (C and D) CD45RA expression in CITE-seq (C) and MERSCOPE (D) dataset. In both datasets, high CD45RA expression level is observed in B cells. Note that based on integration, the CD45RA+ NK cells and naive T cells (arrows) are largely depleted in the MERSCOPE uterine cancer dataset. The same UMAP embeddings are used as (A) and (B). (E and F) CD45RO expression in CITE-seq (C) and MERSCOPE (D) dataset. In both datasets, high CD45RA expression level is observed in T cells. The same UMAP embeddings are used as (A) and (B).
Taken together – the simultaneous profiling of RNA and protein with our MERSCOPE Protein Stain Kit (launched in September of 2022) offers an unparalleled insight into the complex interplay between immune cells and tumor cells, holding significant potential for research uncovering novel biomarkers and therapeutic targets.
There are three ways to explore our multiomic data release:
- Utilize the MERSCOPE® Web Vizualizer to interactively view the full output of the MERSCOPE experiment.
- Download the data files directly to your computer.
- Perform spatial data analysis with Vizgen’s Jupyter Notebook on Google Colab.
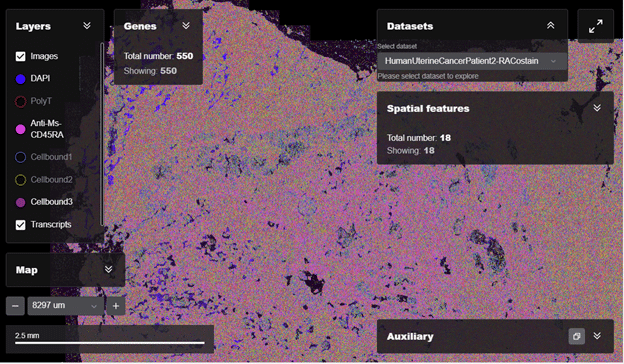
Figure 3: MERSCOPE® Web Vizualizer displaying two MERFISH datasets from human uterine cancer FFPE tissue section generated by the MERSCOPE Platform. The MERSCOPE Web Vizualizer enables users to easily explore the MERFISH datasets. User can zoom from whole-tissue view down to subcellular resolution, visualize the single-cell gene expression of all or a subset of genes, toggle protein stain on and off, color cells based on cell type annotations, and switch between datasets.
“Understanding the interplay between tumor microenvironments and immune cell populations is critical in the development of effective cancer immunotherapies,” said Terry Lo, President and CEO of Vizgen. “At Vizgen, we are proud to continue equipping researchers with the ability to simultaneously profile RNA and proteins within the MERSCOPE Platform to analyze the interactions between tumor and immune cells with the hopes of potentially uncovering novel biomarkers and therapeutics targets.”
Access the dataset today to explore the immuno-oncology landscape of human uterine cancer: https://info.vizgen.com/access-vizgen-data-release-protein-co-detection